Covid-19 and flu: Predicting virus-virus interactions
The interaction between two types of viruses is complex and difficult to predict. Using statistical tests, scientists are trying to decipher the interaction of viruses such as influenza and SARS-CoV-2 from epidemiological data. Researchers at the Max Planck Institute for Infection Biology have now taken a closer look at a test for studying virus-virus interactions. They came to the conclusion that this test design interprets the measure of interaction incorrectly or even in the opposite way. Where viral diseases actually reinforce each other, the test design shows an antagonistic interaction. Reliably predicting virus-virus interactions is an important part of pandemic control—for example, when the COVID-19 pandemic meets the flu season. The results have now been published in the journal Proceedings of the Royal Society.
Catching a viral infection is bad news, but what happens if our body is confronted with two viruses at the same time? Predicting the outcome of co-circulation of viruses is complicated as they can interact both in a synergistic or antagonistic way: Viral infections can inhibit others – this can be due to an already alarmed immune system or viruses “competing” for host cells to infect. But viruses can also amplify each other, for example by making a host more susceptible to infection. And sometimes they simply don’t interact. To know the form of interaction between viruses can be crucial for predicting the course of an epidemic.
Coronavirus meets flu viruses
The research group of Matthieu Domenech de Cellès at the Max Planck Institute for Infection Biology studied co-circulation of viruses with SARS-CoV-2 from the earliest days of the pandemic. They were particularly interested in the interaction between SARS-CoV-2 and influenza viruses. The team modelled the first months of the corona pandemic in Europe, showing that the decrease of Covid-19 cases in spring 2020 was not only related to countermeasures, but also to the end of flu season. They concluded that Influenza and SARS-CoV-2 may interact in a synergistic manner—an interaction that was supported by some experimental studies, but that requires further epidemiological investigations. The results suggested that measures against influenza, like a flu shot, may also help against Covid-19. However, after publishing their findings in a first preprint, other publications came to opposing results and proposed an antagonistic interaction between SARS-CoV-2 and Influenza viruses.
Flu and Covid-19 – with or against each other?
This contradiction gave the team around Domenech de Cellès the impetus to further investigate common models for studying virus-virus interactions at the population scale. Studies that had reported an antagonistic interaction used an epidemiological design called the test-negative design. This design was used to estimate the difference in risk of SARS-CoV-2 infection between individuals testing positive or negative to influenza. Specifically, if influenza and SARS-CoV-2 do not interact and therefore circulate independently, then one expects the frequency of co-detection estimated from cross-sectional data to be approximately equal to the product of each virus’s detection frequency. This intuition originates from standard statistics: for two unrelated events (like rolling two dice), one calculates the joint probability of both events by multiplying the probabilities of each event.
“Misleading measures of interaction”
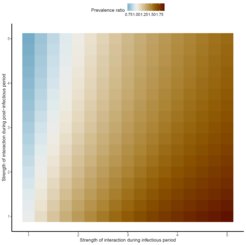
“Although this seems intuitive, we also knew from previous studies, that such simple statistics can be misleading measures of interaction”, states Matthieu Domenech de Cellès in a blog article on his recent publication. To test the epidemiological design, the team created a simulation study—a mathematical model that allowed them to simulate the co-circulation of two respiratory viruses causing seasonal epidemics. With striking results: The studies based on the test-negative design systematically under-estimated the strength of interaction, and could even misclassify interactions that persist after clearance of infection. That way, antagonistic interactions between viruses could be falsely labelled as synergistic. Apparently, respiratory viruses with a high reproductive number and a short duration of infection cannot be met with a straightforward statistical test. Domenech de Cellès concludes: “The complex dynamics of respiratory viruses makes the interpretation of seemingly intuitive measures of interaction difficult, if not impossible”. Other statistical or mathematical models should and must be further employed, as it is becoming clear that corona does not circulate in isolation but within other polymicrobial systems.
