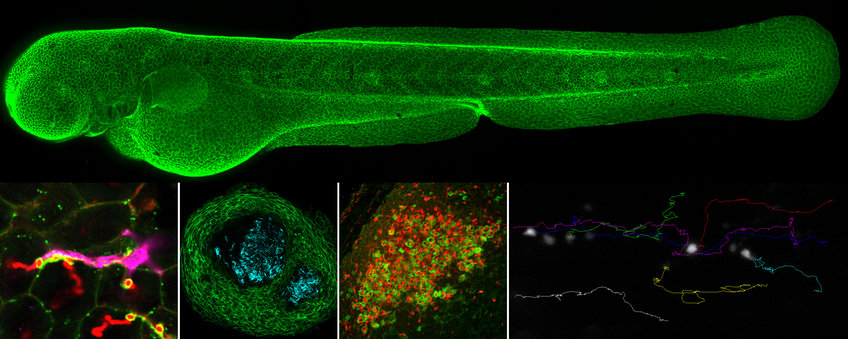
In vivo cell biology of infection
Cell biology of tuberculosis and the granulomatous response
Mycobacterium tuberculosis, the causative agent of tuberculosis, is the leading cause of death in the world by a single infectious agent. We model tuberculosis in the zebrafish using M. marinum infection, a natural pathogen of zebrafish that is a close relative of the M. tuberculosis complex. Optically transparent zebrafish enable us to directly visualize host-bacterial interactions during infection with mycobacteria as well as other bacterial pathogens. Using zebrafish and bacterial genetic tools, cellular reporters and high-resolution microscopy we can study host and bacterial signaling events and cell populations that shape these immune responses. We are particularly focused on understanding the structure and signaling that underlie the granuloma, an aggregate of immune cells that is the central structure of tuberculosis.
Host-directed therapies
Pathogens have evolved many mechanisms to subvert host immune responses and limit pathogen recognition and killing. By understanding immune cell behavior and signaling as well as how pathogens manipulate these responses, we can identify points of therapeutic intervention to facilitate host killing of bacteria and enhance the activity of existing antibacterial therapies. Our ongoing goal is to translate our findings on the immune cell biology of mycobacterial infection and other pathogens into novel host-directed therapies. Additionally, we will take unbiased approaches to discover new therapeutic targets. We have developed an ex vivo mycobacterial granuloma culture platform that will enable chemical and genetic screening within the context of the granuloma. We will also use in vivo chemical genetic screening approaches in zebrafish larvae to identify host-directed therapies to mycobacteria in distinct cellular contexts as well as other bacterial pathogens.