Here to stay: How gut bacteria withstand immune responses during infection
Bacteria of the microbiome can resist antimicrobial peptides of the immune system—the Iatsenko lab now uncovered the mechanism behind this resilience.
Bacterial gut infections? The immune system has an answer for that: broad-spectrum toxins called antimicrobial peptides that kill bacteria and other microorganisms. But some bacteria can resist this challenge. Enter the gut's microbiota, a community of bacteria that often coexist peacefully with their host, aiding digestion and nutrient absorption. During an infection, some of these microbes persist while harmful bacteria are eliminated by the immune system. Now, Igor Iatsenko's research group at the Max Planck Institute for Infection Biology has unraveled the secret to this resilience. Using the fruit fly and a common gut bacterium as a model system for host-microbe interaction in the gut, the researchers showed that modifications in the bacterial cell wall are the key to antimicrobial resistance—a trait conventionally associated with harmful bacteria. These findings shed light on the shared molecular foundations of harmful and non-harmful bacteria in host-microbe interactions. The team's discoveries are now published in the Proceedings of the National Academy of Sciences.
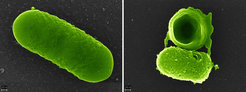
The immune system has many tools to get rid of harmful bacteria. Some are very specific, such as antibodies, while others paint with a broad brush. The latter include antimicrobial peptides, which eliminate harmful bacteria of all kinds by disrupting their cell membranes. However, there is a problem with targeting bacteria in general. Humans, like most other multicellular organisms, are inhabited by microorganisms that are not harmful, but help with digestion and sometimes even prevent infection. Fortunately, our immune system does not eliminate all bacteria: The microbiome survives most infections and its bacteria are not as affected by antimicrobial peptides as harmful bacteria.
The fruit fly’s gut as a model system
In their latest study, the Iatsenko Lab at the Max-Planck-Institute for Infection Biology set out to investigate this resilience. The team around group leader Igor Iatsenko wanted to find out why some bacteria of the microbiome (also known as microbiota) are resistant to antimicrobial peptides during infection. To study the interplay between the host, its microbiome and harmful bacteria, the team employed a model system: the common fruit fly, Drosophila melanogaster. The fly’s gut is much simpler than that of humans, but in principle it works the same and most importantly it is also inhabited by several species of microbiota.
When PhD student Aranzazu Arias-Rojas, first author of the new study, tackled this question in the lab, she first searched for a bacterium that showed significant resistances against antimicrobial peptides. To do this, she infected several hundred fruit flies under varying conditions to see which bacteria remained in the flies after infection. The gut bacterium Lactiplantibacillus plantarum (L. plantarum in short) was a hit. It was still present in most of the flies after infection, suggesting that it was not affected by antimicrobial peptides.
Using ‘jumping genes‘ to find resilience factors
After this first step, the researchers wanted to find the genes responsible for the resistance in order to uncover the underlying molecular mechanism. A transposon screen was used to identify these genes. Transposons are known as jumping genes, because of their ability to change their position in the genome. In a transposon screen, this property is used to knock out genes: When a transposon is introduced into a bacterium, it inserts itself randomly in the DNA, blocking the gene at that location.
The researchers created around 5000 transposon mutants of L. plantarum which were then cultured with polymyxin, an antibiotic that works like antimicrobial peptides. Mutants that failed to grow in the presence of polymyxin had lost their resistance against antimicrobial peptides due to a transposon-deactivated gene. By checking where the transposon had been inserted, the researchers were able to identify the deactivated genes responsible for resistance. In total, the team found three blocked genes that impaired the resistance, among them one gene for cell wall modifications, the dlt operon.
‘Bad’ and ‘good’ bacteria—not so dissimilar after all
The mechanism controlled by this gene was already known—not in microbiota, but in harmful bacteria. Cell wall modifications make harmful bacteria more dangerous by increasing their resistance against antimicrobial peptides. “We did not expect to find this resistance mechanism in microbiota, but it made sense given previous evidence that ‘bad’ and ‘good’ bacteria are not so different after all”, says Arias-Rojas. These shared molecular mechanisms raise questions: what makes a bacterium harmful, and how can microbiota switch to a harmful behavior in their host? Following their work with fruit flies and L. plantarum, the researchers are already pursuing these questions in current projects.
The differences and similarities between harmful bacteria and microbiota also play a role in medicine. Researchers are currently investigating antimicrobial peptides as therapeutics. Antibiotics—today’s standard treatment against bacterial infection—are very efficient in eliminating all bacteria, including our microbiome. This often causes side effects like digestive problems, especially when the treatment is carried out for a long time. As Iatsenko’s study shows, microbiota have adapted to antimicrobial peptides to survive in their host. In the future, antimicrobial peptides could possibly be used as therapeutics, clearing harmful bacteria while leaving the microbiota intact.
